One stretching cell¶
The example in examples/stretchCell presents a validation example for the
material models used in HemoCell. A single cell is initialised in a
periodic domain, i.e. all external surfaces are subjected to periodic boundary
conditions.
The setup mimics the optical-tweezer stretching measurement. A pair of external forces are applied on the outer points of the cell pointing in opposite directions. These forces stretch the cell, where its deformation can be validated with respect to experimental data.
After compilation, the example can be run using single core as:
# run the simulation from the `examples/stretchCell/`
mpirun -n 1 ./stretchCell config.xml
# generate Paraview compatible output files
../../scripts/batchPostProcess.sh
Note
The stretching cell examples should be run with just a single processor, so
either mpirun -n 1 ./stretchCell config.xml as above, or directly
./stretchCell config.xml, as the helper function used to specify the stretching forces on the cell only
supports sequential operation.
The outcome files are generated in tmp/, where the flow field and particle
fields can be visualised separately by viewing the tmp/Fluid.*.xmf and
tmp/RBC.*.xmf files in Paraview.
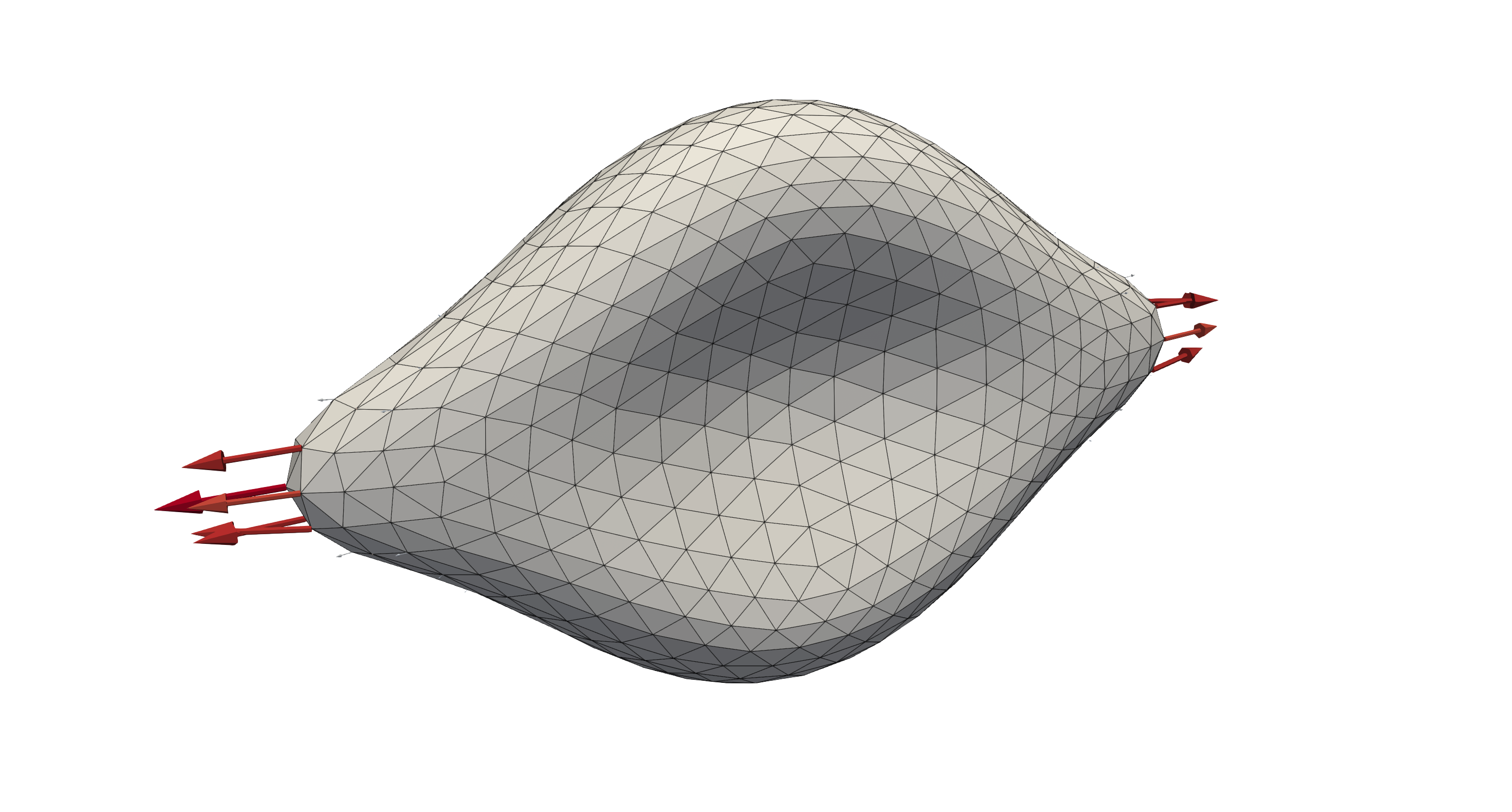
A deformed red blood cell after being subjected to opposing forces at either side of the RBC, where the arrows visualise the applied forcing.¶
Note
The stretchCell executable also collects the axial and transverse
dimensions of the cell for each iteration for the used stretch force $f.
These are collection in ./stretch-$f.log and can be used for validation
purposes.
Configuration¶
There is one parameter that can be modified in this example, i.e. the applied stretching force in pico Newton:
<parameters><stretchForce>: the applied stretching force in pico Newton.
Validation¶
The script validation.sh helps to perform basic validation of the cell
stretch model as originally presented in [1]. The script runs
through various evaluations of stretchCell with different values for the
considered stretch force stretchFoce (Configuration). The resulting
log files stretch-*log contain the axial and transverse dimensions of the
RBC for the different stretch forces. From here, Figure 4
([1]) can be recreated. If gnuplot is available on the
system, the figure is automatically generated under
stretchCell/validation/validation.png.
After compilation stretchCell the validation can be
evaluated through:
./validation.sh
open validation/validation.png
Note
In validation/ a number of reference values are provided that correspond
to the curves presented in [1] which provide a clear
reference point when investigating cell stretch models.
